Molecules Molecules Molecules!
This week, I’ve been engrossed in making molecular movie magic! When I was a student at UIC, we learned about the Protein Data Bank, and I thought that was pretty cool. We would bring files into a free program called Chimera, clean them up a bit, and export them as .stl or .obj files, anything we could pull into 3DsMax. I think that with larger files, you would also have to take the intermediate step of going into Mesh Lab to try and reduce your polygon count. From there, you could put any kind of material on them, set up lighting, do anything you wanted to.
I think that this was the first one I ever tried…
The image itself may not be all that impressive on it’s own right, but I remember being excited because I’d found something related to a good friend’s graduate work. I wish I could remember the name of it now, but I know it was tied to the magnetic bacteria that Cody studied when he was at CalTech. And just the thought of being able to search for and find 3D data related to a friend’s work like that, to be able to bring it into a 3D space and look at it, and then further still, be able to make something of it, and render it into an image that could be shared, well that all seemed pretty exciting.
A year or two later, they came out with Molecular Maya. This took out all the difficulty of working with Chimera, and allowed you to pull data directly from the PDB (Protein Data Bank) into Maya (which is another 3D animation program, much like 3DsMax).
Since coming to work for Sapling Learning, I have been using Cinema 4D (as opposed to either Maya or 3DsMax). And I have a plug-in downloaded called ePMV which is short for Embedded Python Molecule Viewer. It’s pretty awesome. Much like with Molecular Maya, you get to bring files directly in to your 3D scenes, without having to use other software in the interim.
I’m not sure if it didn’t exist before, or if I just didn’t know about it my first times trying to navigate the PDB, but finding PDB 101 was been a great help in being able to find the right proteins when looking for something specific. It’s also been a wealth of information when I’m trying to understand more about a molecule that I’m working with.
But I have to say that what I’m really excited about this week, is the realization that Marvin Sketch can be used to get a .pdb file on any molecule that you can draw. A coworker/chemistry genius showed me the Marvin Sketch Molecule Viewer, and I knew it was awesome when he did, but the more I get my head around all this, the more awesome this tool becomes. You can literally draw out any molecule you know the arrangement for, ask it to clean it up in either 2D or 3D for you, and if you’re looking at the 3D, you can export the file as a .pdb and just like that you have everything you need to bring it in via ePMV and use it for static illustration or animation.
Now this still leaves me limited by the fact that I really don’t have a background in chemistry, and I often don’t know how to draw out the thing that I want (though I am learning a lot of that sort of stuff these days). So how amazing did I feel when just this week I noticed SMILES (Simplified molecular-input line-entry system.) Oh yeah. So there I am, scouring the internet for a clean model of ATP (adenosine triphosphate). And there are literally thousands of references to it within the PDB but it’s always with something. I wanted to find it by itself. And then I’m looking at the Wikipedia page on it. They have 3D models featured. Clearly this data is out there. I should be able to find it. And then I’m looking at the formula for ATP, and it’s C10H16N5O13P3. So I think to myself, that if I were better at drawing out molecules, I could just get it out of Marvin sketch. But sometimes Wikipedia has links to PDB pages, where was that? Well it didn’t have that but it did have a listing where you could show SMILES data
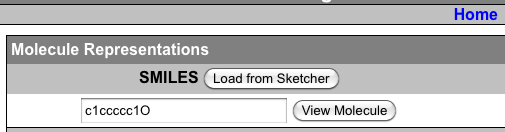
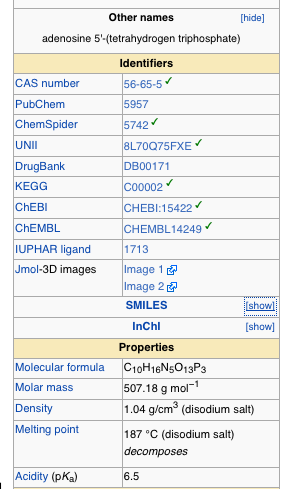 And if you just click the box that says show, next to SMILES, it gives you a code. You copy that code and paste it right into Marvin Sketch Molecule View SMILES loader
And if you just click the box that says show, next to SMILES, it gives you a code. You copy that code and paste it right into Marvin Sketch Molecule View SMILES loader
And just like that, you can get any molecule that you have that code for. I did this for ATP, ADP, glucose, and glucose 6 phosphate. They all had a SMILES code, listed right there in Wikipedia. And I was able to save .pdb files right my local computer’s desktop. And when I pulled them all in to Cinema 4D with the ePMV plug-in, I realized that it was easy to see exactly what atoms are coming off of one molecule and added to the other in the process I am working with. And because it was all so accessible like that, we’re going to be able to show people that, really clearly.
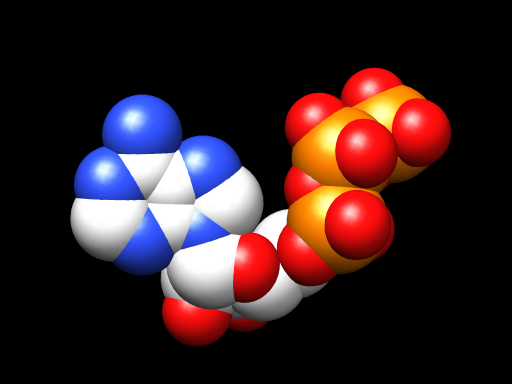
I don’t have any of my files from work handy here, but just now I put ATP’s code into the SMILES loader of Marvin Sketch and exported a 3D PDB file. I opened the file using Chimera and in less than a minute had this.
And if I were doing this in a 3D animation program, I could take things further still and make it pretty, make it move. Or I could grab that last bundle of spheres there, the last yellow (phosphorous) and the reds (oxygen atoms) clinging to it, and pop that right off to show you the difference between ATP and ADP, and what exactly that means when people talk about that happening.
I think that’s really friggin cool!
Well, I suppose I should stipulate that we’re *almost* seeing exactly how that works. I just learned today that there’s hydrogen in there too, but chemists seem to have this game of hide-the-hydrogen that they like to play on the rest of us. They claim it’s clearer that way. They’re probably right, but I’m not 100% sold yet. Still, except for the invisible hydrogen atoms, this is what ATP looks like. This is it’s shape. And all those reactions in the body where ATP gets converted to ADP are just this little guy getting a little bit shorter when that last phosphate group teams up somewhere else.
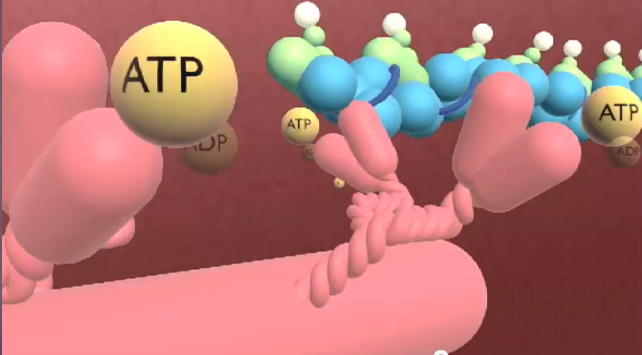
When I made my sliding filament animation, I remember having no idea what ATP actually looked like. I remember talking to other people about it, and none of the people I spoke to knew either. I wound up representing it with big yellow spheres in the animation. And that worked for the piece that it was, and was actually very clear in terms of what needed to be shown there. But I just love that this week, while trying to figure something out about ATP I wound up with not only a better picture of it, but my own model to play with and animate.
I suppose that lately it’s just been molecules molecules molecules for me. I never thought I would get this into the micro world, but I have to say, I think it was time. And with all this technology, all these programs making it easier than ever to really see and get a handle on what everything really is, well, I can’t think of a better time to be learning this stuff.